The dataset provided here aims at predicting protein disorder (continuous). The disorder values used here are CheZOD scores, where values below 8 signify disorder and values above 8 signify order.
The provided dataset is compiled as follows:
- Training, Test: The SETH dataset was used and double-checked. Three duplicated sequences were excluded from the training set (see below).
- Validation: Random 0.1 split from training set
Removed Sequences
There were three duplicates in the training set with ambiguous score annotations that were removed:The dataset is provided in biotrainer-ready fasta format. Each entry contains a sequence and a header, providing the sequence id, the set (train/val/test), the target scores (separated by ';') and masks (unresolved (0): disorder value == 999, otherwise resolved (1)).
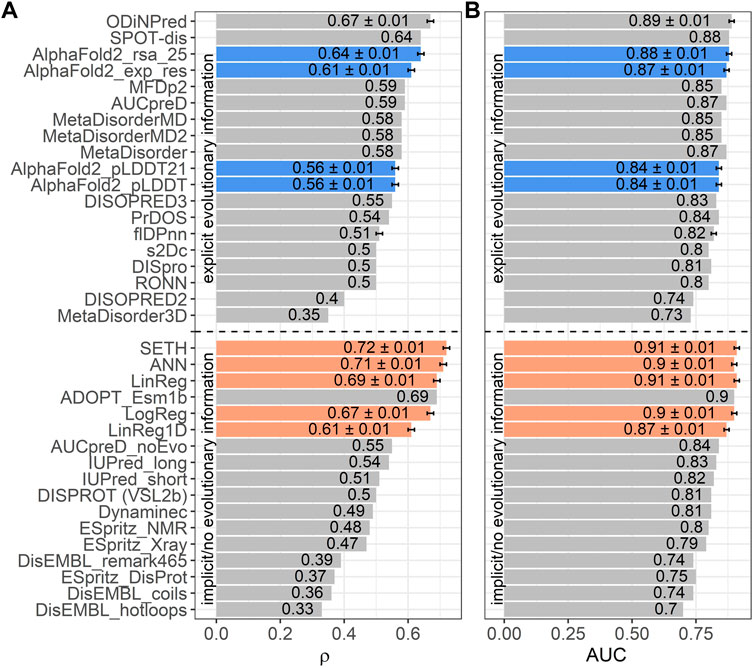
The SETH paper provides benchmarks for the disorder prediction task on an almost identical dataset (training set - 3 duplicates, identical test set).
@InBook{Nielsen2020,
author = {Nielsen, Jakob T. and Mulder, Frans A. A.},
pages = {303--317},
publisher = {Springer US},
title = {Quantitative Protein Disorder Assessment Using NMR Chemical Shifts},
year = {2020},
isbn = {9781071605240},
booktitle = {Intrinsically Disordered Proteins},
doi = {10.1007/978-1-0716-0524-0_15},
issn = {1940-6029},
}@Article{Nielsen2019,
author = {Nielsen, Jakob T. and Mulder, Frans A. A.},
journal = {Scientific Reports},
title = {Quality and bias of protein disorder predictors},
year = {2019},
issn = {2045-2322},
month = mar,
number = {1},
volume = {9},
doi = {10.1038/s41598-019-41644-w},
publisher = {Springer Science and Business Media LLC},
}@Article{Ilzhoefer2022,
author = {Ilzhöfer, Dagmar and Heinzinger, Michael and Rost, Burkhard},
journal = {Frontiers in Bioinformatics},
title = {SETH predicts nuances of residue disorder from protein embeddings},
year = {2022},
issn = {2673-7647},
volume = {Volume 2 - 2022},
abstract = {Predictions for millions of protein three-dimensional structures are only a few clicks away since the release of AlphaFold2 results for UniProt. However, many proteins have so-called intrinsically disordered regions (IDRs) that do not adopt unique structures in isolation. These IDRs are associated with several diseases, including Alzheimer’s Disease. We showed that three recent disorder measures of AlphaFold2 predictions (pLDDT, “experimentally resolved” prediction and “relative solvent accessibility”) correlated to some extent with IDRs. However, expert methods predict IDRs more reliably by combining complex machine learning models with expert-crafted input features and evolutionary information from multiple sequence alignments (MSAs). MSAs are not always available, especially for IDRs, and are computationally expensive to generate, limiting the scalability of the associated tools. Here, we present the novel method SETH that predicts residue disorder from embeddings generated by the protein Language Model ProtT5, which explicitly only uses single sequences as input. Thereby, our method, relying on a relatively shallow convolutional neural network, outperformed much more complex solutions while being much faster, allowing to create predictions for the human proteome in about one hour on a consumer-grade PC with one NVIDIA GeForce RTX 3060. Trained on a continuous disorder scale (CheZOD scores), our method captured subtle variations in disorder, thereby providing important information beyond the binary classification of most methods. High performance paired with speed revealed that SETH’s nuanced disorder predictions for entire proteomes capture aspects of the evolution of organisms. Additionally, SETH could also be used to filter out regions or proteins with probable low-quality AlphaFold2 3D structures to prioritize running the compute-intensive predictions for large data sets. SETH is freely publicly available at: https://github.com/Rostlab/SETH.},
doi = {10.3389/fbinf.2022.1019597},
url = {https://www.frontiersin.org/journals/bioinformatics/articles/10.3389/fbinf.2022.1019597},
}@Article{Dass2020,
author = {Dass, Rupashree and Mulder, Frans A. A. and Nielsen, Jakob Toudahl},
journal = {Scientific Reports},
title = {ODiNPred: comprehensive prediction of protein order and disorder},
year = {2020},
issn = {2045-2322},
month = sep,
number = {1},
volume = {10},
doi = {10.1038/s41598-020-71716-1},
publisher = {Springer Science and Business Media LLC},
}The RAW data downloaded from the aforementioned publications is subject to GPL v3.0. Modified data available in this repository falls under AFL-3.